Recently, NVIDIA CEO Jensen Huang announced updates to the open beta of NVIDIA Merlin, an end-to-end framework that democratizes the development of large-scale deep learning recommenders. With NVIDIA Merlin, data scientists, machine learning engineers, and researchers can accelerate their entire workflow pipeline from ingesting and training to deploying GPU-accelerated recommenders (Figure 1).
While NVIDIA Merlin provides end-to-end components, this post focuses on the latest updates to Merlin NVTabular, the extract, transform, load (ETL) component of Merlin. For more information about the other components, see Announcing NVIDIA Merlin: An Application Framework for Deep Recommender Systems.
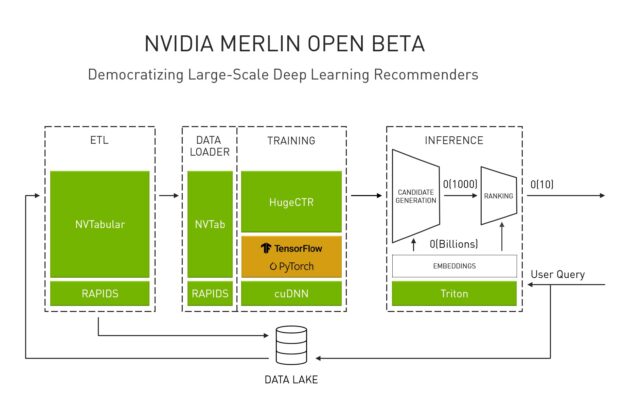
NVTabular is the ETL component of Merlin, introduced to address the common pain points of the data pipeline for recommender systems. When training industrial recommender systems, it is common to see training datasets of multi-terabyte, or even petabyte scale, comprising billions of user-item interactions. Preprocessing or doing feature engineering for datasets of this size is an extremely time consuming task, so much so that there’s a common adage within the data science community that data scientists spend more of their time on ETL and data preparation than model training and fine-tuning.
The initial release of Merlin brought the basic functionality of ETL for recommender systems to the GPU. In this open beta release we have added multi-GPU support through a rework of the entire backend of NVTabular. NVTabular now uses Dask and Dask-cuDF which greatly improves interoperability with the RAPIDS ecosystem. While we had initially built our own custom iterator for dealing with datasets larger than GPU memory, Dask-cuDF proved much more efficient, even on a single GPU, reducing ETL runtimes on the Criteo Terabyte Click Log dataset from 22 mins to 14 mins on V100. This allowed us to easily scale across multiple GPUs on a single node. This backend change also unified the single and multiple GPU versions making code development easier, both for us and for anyone developing custom ops.
During the development for this release, we studied the data loading of the major frameworks and developed new ways to get tabular data efficiently onto the GPU. If not optimized and accelerated, loading huge amounts of data into deep learning (DL) frameworks can become a common bottleneck resulting in substantial underutilization of the GPUs while training. We developed custom data loaders to improve the performance of training recommender systems in TensorFlow and PyTorch. We will be working across both ecosystems to integrate these techniques into other data loading libraries so that they are easy to use. Our techniques to optimize the training were built from existing work by the frameworks and provide an optimal way of loading data onto the GPU. In the future, we hope to make them a part of the frameworks.
We also worked to integrate these data loader optimizations directly into HugeCTR, the recommender system specific training framework of Merlin. HugeCTR has been specifically designed for training DL recommender systems and has its own GPU optimized data loader. In the Merlin open beta release, HugeCTR has added a new Parquet data reader to digest NVTabular preprocessed data more easily and efficiently. We will continue to expand that integration.
Finally, we have added a significant number of new operations to NVTabular, summarized in this post.
Multi-GPU support using RAPIDS cuDF and Dask
In this latest release, NVTabular supports multi-GPU with RAPIDS cuDF, Dask, and Dask_cuDF.
- cuDF—A Python GPU DataFrame library built on the Apache Arrow columnar memory format for loading, joining, aggregating, filtering, and otherwise manipulating tabular data using a DataFrame-style API.
- Dask—A flexible library for parallel computing in Python that makes scaling out your workflow smooth and simple. On the CPU, Dask uses pandas to execute operations in parallel on DataFrame partitions.
- Dask-cuDF—Extends Dask where necessary to allow its DataFrame partitions to be processed by cuDF GPU DataFrames as opposed to pandas DataFrames. For instance, when you call
dask_cudf.read_csv, your cluster’s GPUs do the work of parsing the CSV files with the underlyingcudf.read_csvcall.
Under the hood, NVTabular uses all these technologies to provide a high-performance, recommender system-specific, ETL pipeline on multiple GPUs.
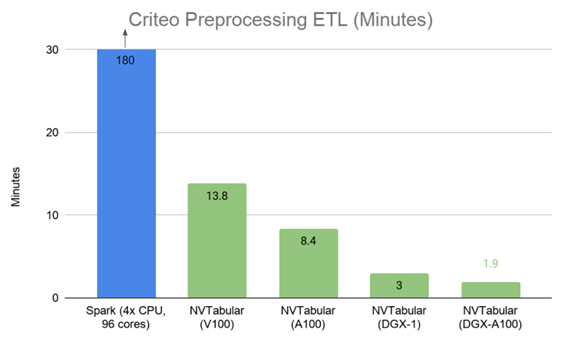
Figure 2 shows a 95x speedup using NVTabular multi-GPU on the DGX A100 compared to Spark on a four-node, 96 vCPU core, CPU cluster processing 1.3 TB of data in the Criteo Terabyte dataset, providing a speedup of 5.3x using eight A100 GPUs, from 10 mins on 1xA100 to 1.9 mins on 8xA100.
Here’s the process of using NVTabular on multiple GPUs. In the beginning of the workflow, you initialize a LocalCUDACluster instance with multiple GPUs. NVTabular uses the environment to execute the workflow on the defined GPUs. In this post, we show a shortened example but you can get the full multi-gpu_dask.ipynb tutorial in the GitHub repo.
Initialize the multi-GPU cluster and define the protocol (TCP/IP or UCX), GPU IDs, and host memory. Then, create a client as a local CUDA cluster. Because allocating memory is often a performance bottleneck, we recommend initializing a memory pool on each worker. When using a distributed cluster, you must use the client.run utility to make sure that a function is executed on all available workers, using _rmm_pool.
# Initialize the local multi-GPU cluster protocol = "tcp" # "tcp" or "ucx" visible_devices = "0,1,2,3,4,5,6,7" # Select devices to place workers device_memory_limit = "28GB" # Spill device mem to host at this limit memory_limit = "96GB" # Spill host mem to disk near this limit cluster = None # (Optional) Specify the existing scheduler port if cluster is None: cluster = LocalCUDACluster( protocol = protocol, CUDA_VISIBLE_DEVICES = visible_devices, local_directory = dask_workdir, device_memory_limit = parse_bytes(device_memory_limit), memory_limit = parse_bytes(memory_limit), ) # Create the distributed client client = Client(cluster) # Initialize RMM pool on ALL workers def _rmm_pool(): cudf.set_allocator( pool=True, initial_pool_size=None, # Use default size allocator="default", ) client.run(_rmm_pool)
After the local CUDA cluster is defined, you can define and execute the NVTabular workflow.
# Initialize the workflow workflow = nvt.Workflow(cat_names=cat_names, cont_names=cont_names, label_name=label_name, client=client) # Add continuous operations workflow.add_preprocess(ops.Normalize(columns=["x", "y"])) # Add categorical operations workflow.add_preprocess( ops.Categorify( out_path=demo_output_path, # Path to write unique values used for encoding ) ) # Finalize the workflow workflow.finalize() workflow.apply( dataset, output_format="parquet", output_path=os.path.join(demo_output_path,"processed"), shuffle=Shuffle.PER_WORKER, out_files_per_proc=8, )
Data loading for recommender systems
The initial data loaders for the deep learning frameworks were designed for loading images. The large chunks of data were gathered, item by item, and collated into a batch to be passed to the GPU for training. For the tabular data that you commonly see in recommender systems, the amount of data per example is trivial. If you select item by item, particularly across a huge dataset of terabytes in scale, data batch creation is slow. At training time, the GPU is never fully utilized and that training on GPU isn’t any faster than on CPU.
To break this paradigm, we used an iterable data loader, an idea introduced by the PyTorch team. By treating the data loader as an iterator, you can pass whole batches directly to the training framework much more efficiently than if you’re creating batches item by item. This raises some additional constraints in terms of how the data is shuffled. NVTabular data loaders use two stages of shuffling to ensure that data in a batch is random per epoch.
At a high level, you are shuffling the data during feature engineering, again during data loading. You create a buffer of data on the GPU that is close to uniformly random. However, that data can be easily shuffled in the next epoch to create a new random order without requiring a full shuffle of the entire dataset. This buffer is loaded asynchronously while the training happens within the framework.
The current data loader supports only single, hot, categorical variables and continuous. Multi-hot support and session-based support is coming soon. With TensorFlow, you use the same core concept of iterators. TensorFlow already supports this and provides a few additional helper embedding layers to efficiently accelerate the access of data on the GPU. Both the TensorFlow and PyTorch data loaders share the same backend code to create the iterator. They have framework-specific wrappers to format the data correctly for the corresponding framework. Using dlpack passes the data between the cuDF representation and the frameworks efficiently, significantly reducing overhead.
In preliminary testing, we see a significant order of magnitude speedup over the item-by-item data loading mechanisms that are more commonly used by practitioners in both frameworks. Even more excitingly, we see high GPU utilization and a further 2-5x speedup from automatic mixed precision (AMP). which is not seen in item-by-item data loading because the GPU is not utilized. We’ll have much more to share in our 0.3 release of NVTabular. We wanted to share these exciting results and get your feedback as you try them out.
An example of acceleration in TensorFlow
The NVTabular data loader for TensorFlow is designed to feed tabular data efficiently to deep learning model training with TensorFlow. In this post, we discuss the main component but the NVTabular demo on Rossmann data provides a detailed example. For more information, see the Jupyter notebook.
Here’s a concrete example. First, you must define some basic parameters, such as the training and validation data, data schema, which columns are categorical and numerical, and the embedding output dimensions with nvt.ops.get_embedding_sizes. For more information, see the demo on Rossmann data.
Afterwards, you can initialize the NVTabular data loader for TensorFlow, using the following code example. Before the import, it’s important to define the environment variables for controlling the TensorFlow memory usage. After the import, you create native TensorFlow input features for categorical columns (tf.feature_column.embedding_column) and numerical columns (tf.feature_column.numeric_column). Use these to initialize the new Merlin NVTabular data loader KerasSequenceLoader.
import tensorflow as tf # Control how much memory to give TensorFlow with this environment variable # IMPORTANT: Do this before you initialize the TensorFlow runtime, otherwise # it's too late and TensorFlow will claim all free GPU memory os.environ['TF_MEMORY_ALLOCATION'] = "8192" # explicit MB os.environ['TF_MEMORY_ALLOCATION'] = "0.5" # fraction of free memory from nvtabular.loader.tensorflow import KerasSequenceLoader, KerasSequenceValidater # wrapper to keep things tidy def make_categorical_embedding_column(name, dictionary_size, embedding_dim): return tf.feature_column.embedding_column( tf.feature_column.categorical_column_with_identity(name, dictionary_size), embedding_dim ) # instantiate the columns categorical_columns = [ make_categorical_embedding_column(name, *EMBEDDING_TABLE_SHAPES[name]) for name in CATEGORICAL_COLUMNS ] continuous_columns = [ tf.feature_column.numeric_column(name, (1,)) for name in CONTINUOUS_COLUMNS ] # feed them to the datasets train_dataset_tf = KerasSequenceLoader( TRAIN_PATHS, # you could also use a glob pattern feature_columns=categorical_columns+continuous_columns, batch_size=BATCH_SIZE, label_names=LABEL_COLUMNS, shuffle=True, buffer_size=0.06 # amount of data, as a fraction of GPU memory, to load at one time ) valid_dataset_tf = KerasSequenceLoader( VALID_PATHS, # you could also use a glob pattern feature_columns=categorical_columns+continuous_columns, batch_size=BATCH_SIZE*4, label_names=LABEL_COLUMNS, shuffle=False, buffer_size=0.06 # amount of data, as a fraction of GPU memory, to load at one time )
You can then define and initialize the model with TensorFlow with no additional changes required. Finally, call the model.fit function with the optimized KerasSequenceLoader.
validation_callback = KerasSequenceValidater(valid_dataset_tf) history = tf_model.fit( train_dataset_tf, callbacks=[validation_callback], epochs=EPOCHS, )
An example of acceleration in PyTorch
NVTabular similarly provides an optimized data loader for PyTorch.
This example demonstrates training a deep learning model based on the Rossmann Store Sales dataset from Kaggle, which has data similar to what you see in recommender systems. It is also available as a Jupyter notebook in the GitHub repo.
First, you must define the data schema, the column names for categorical, continuous, and label columns. Next, initialize the Merlin NVTabular PyTorch data loader with the TRAIN_PATHS and VALID_PATHS values containing the filenames of the training and validation datasets.
# TensorItrDataset returns a single batch of x_cat, x_cont, y. collate_fn = lambda x: x train_dataset = TorchAsyncItr(nvt.Dataset(TRAIN_PATHS), batch_size=BATCH_SIZE, cats=CATEGORICAL_COLUMNS, conts=CONTINUOUS_COLUMNS, labels=LABEL_COLUMNS) train_loader = DLDataLoader(train_dataset, batch_size=None, collate_fn=collate_fn, pin_memory=False, num_workers=0) valid_dataset = TorchAsyncItr(nvt.Dataset(VALID_PATHS), batch_size=BATCH_SIZE, cats=CATEGORICAL_COLUMNS, conts=CONTINUOUS_COLUMNS, labels=LABEL_COLUMNS) valid_loader = DLDataLoader(valid_dataset, batch_size=None, collate_fn=collate_fn, pin_memory=False, num_workers=0)
Finally, after you define the model using PyTorch (no changes required), you can use the NVTabular data loader in the training loop.
def process_epoch(dataloader, model, train=False, optimizer=None): model.train(mode=train) with torch.set_grad_enabled(train): y_list, y_pred_list = [], [] for x_cat, x_cont, y in iter(dataloader): y_list.append(y.detach()) y_pred = model(x_cat, x_cont) y_pred_list.append(y_pred.detach()) loss = loss_func(y_pred, y) if train: optimizer.zero_grad() loss.backward() optimizer.step() y = torch.cat(y_list) y_pred = torch.cat(y_pred_list) epoch_loss = loss_func(y_pred, y).item() epoch_rmspe = rmspe_func(y_pred, y).item() return epoch_loss, epoch_rmspe for epoch in range(EPOCHS): train_loss, train_rmspe = process_epoch(train_loader, model, train=True, optimizer=optimizer) valid_loss, valid_rmspe = process_epoch(valid_loader, model, train=False)
New operator support
This release, we worked on a number of workflows with a few customers who helped define the operators added to NVTabular. Of particular note are the operators added to support the winning entry of our RecSys challenge submission. In the example workflow, we provide the same feature engineering workflow that was used by the RecSys and KGMON teams to win the competition.
The following new operators are a part of this release:
Column SimilarityDropnaFilterFillMedianHashBucketJoinGroupbyJoinExternalLambdaOpTargetEncodingDifferenceLag
Share your feedback
As always, we love feedback and suggestions. Join the NVIDIA/NVTabular GitHub repo and share any challenges or ideas about further improvements to make Merlin better suit your needs. We’re actively seeking project contributors. Contact us if your team is interested in using Merlin as a part of a recommender system framework.
