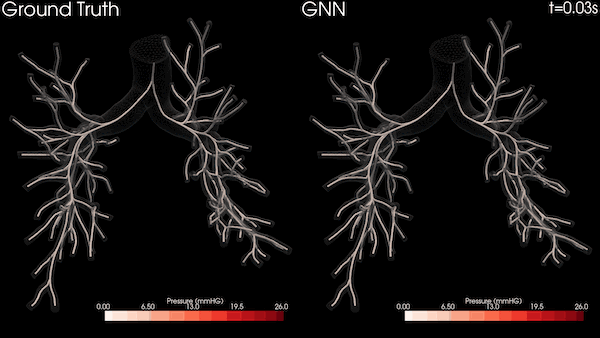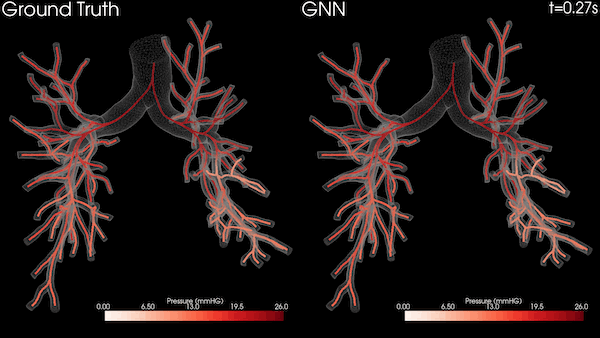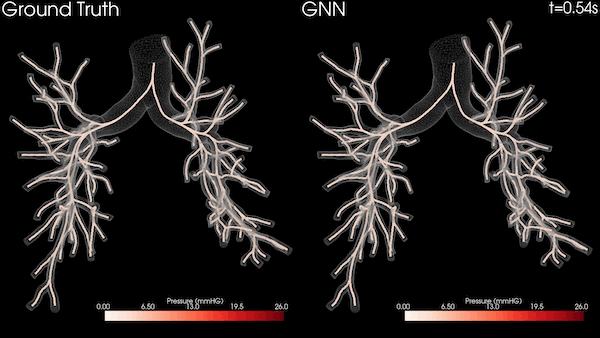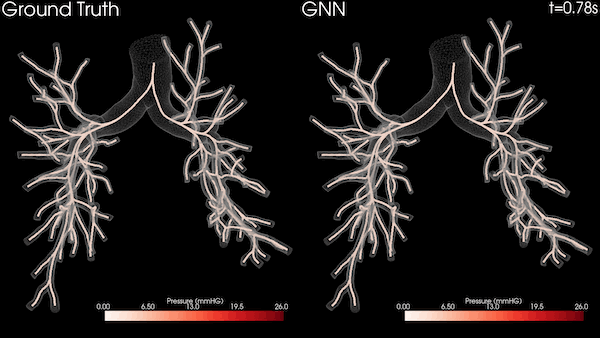
A Stanford University team is transforming heart healthcare with near real-time cardiovascular simulations driven by the power of AI. Harnessing physics-informed machine learning surrogate models, the researchers are generating accurate and patient-specific blood flow visualizations for a non-invasive window into cardiac studies. The technology has far-reaching scope, from evaluating coronary artery aneurysms to pioneering new surgical methods for congenital heart disease and enhancing medical device efficacy. With enormous potential in advancing cardiovascular medicine, the work could offer innovative methods for combating the leading cause of death in the US.
Cardiovascular simulations are important enablers for patient-specific treatment for several heart-related ailments. 3D computational fluid dynamics (CFD) simulations of the blood flow using finite-element methods are a computationally challenging task, especially in clinical practice. As an alternative, physics-based reduced-order models (ROMs) are often employed due to their increased efficiency.
However, such ROMs rely on simplified assumptions of vessel geometry, complexity, or simplified mathematical models. They often fail to model the quantities of interest, such as pressure losses at vascular junctions accurately, which would otherwise require full 3D simulations. Conventional data-driven reduced-order approaches, such as projection-based methods, don’t offer much flexibility with respect to changes to the domain geometry, which is crucial for patient-specific cardiovascular simulations.
As an alternative, employing deep learning-based surrogates can model these complex physical processes. Physics-informed machine learning (physics-ML) enables training deep learning models that can offer both computational efficiency as well as flexibility, through parameterizable models. Recently, graph neural network (GNN) based architectures have been proposed to build physics-ML models for emulating mesh-based simulations. This approach offers generalization over different meshes, boundary conditions, and other input parameters, which makes it a perfect candidate for patient-specific cardiovascular simulations.
The research team from Stanford University leveraged MeshGraphNet, a graph neural network (GNN)-based architecture, to devise a one-dimensional Reduced Order Model (1D ROM) for simulating blood flow. The team implemented this approach into NVIDIA PhysicsNeMo, a platform equipped with an optimized implementation of MeshGraphNet. The reference MeshGraphnet implementation in NVIDIA PhysicsNeMo brings several code optimizations such as data parallelism, model parallelism, gradient checkpointing, cuGraphs, and multi-GPU and multi-node training. All of which are useful for the development of GNNs for cardiovascular simulation.
NVIDIA PhysicsNeMo is an open-source framework geared towards the development of physics-informed machine learning models. It enhances the efficacy of high-fidelity, complex multi-physics simulations by enabling the exploration and development of surrogate models. This framework enables seamless integration of datasets with first principles, whether described by the governing partial differential equations or other system attributes, such as physical geometry and boundary conditions. Furthermore, it provides the capability to parameterize the input space, fostering the development of parameterized surrogate models critical for applications like digital twins.
NVIDIA PhysicsNeMo provides a variety of reference applications across domains from CFD, thermal, structural, and electromagnetic simulations, and can be used for numerous industry solutions ranging across climate modeling, manufacturing, automotive, healthcare, and more. These examples can serve as a foundation for the work of researchers, such as vortex shedding, in this case for the Stanford research team.
Modeling cardiovascular simulation as an AI problem
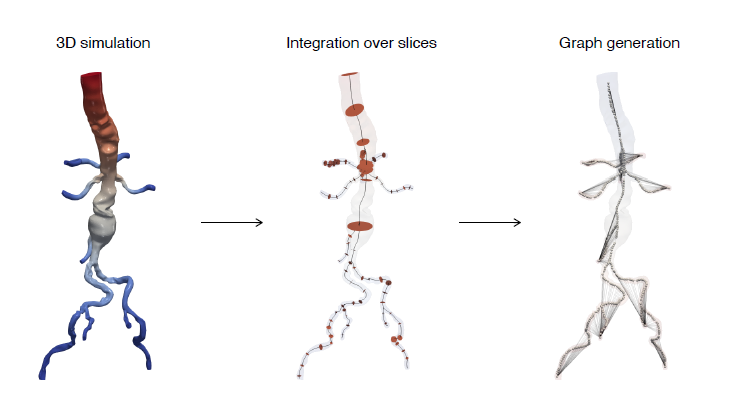
In this research study, the Stanford team’s objective was to develop a one-dimensional physics-ML model. They chose GNNs for developing a surrogate that infers the pressure and flow rate along the centerline of compliant vessels. The team used geometry from 3D vascular models and generated a directed graph consisting of a set of nodes along the centerline of the geometry.
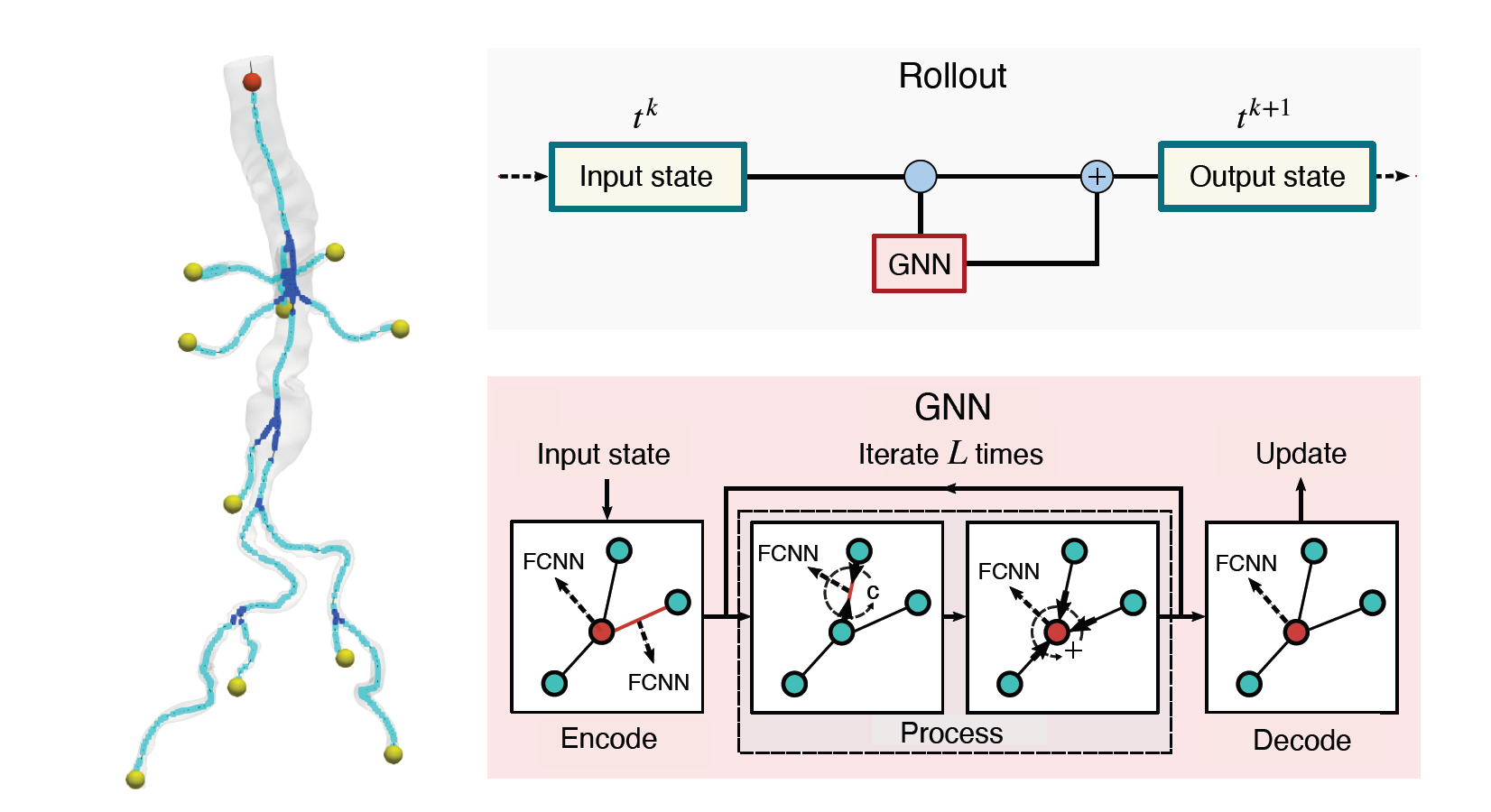
The nodes and edges of the graph capture the state of the system at a given time. For instance, the cross-sectional average pressure, flow rate, and area of the vessel’s lumen along the centerlines of the vessel are included as node features. The GNN takes the state of the system at time t and infers the state of the system at the next time step. This can be iteratively applied to a rollout for simulating the cardiovascular cycle as shown in Figure 2.
The boundary conditions at the inlet (red) and outlet (yellow) are required for determining the hemodynamics in the vessels. These are modeled as special edges of the graph to account for the effect and the complexity of these boundary conditions. Boundary condition parameters, one-hot vector encoding for differentiating between nodes (branch, junction, inlet, outlet), and minimum diastolic and maximum systolic pressure in the cardiac cycle are also included as node features.
For the details on the modeling of the physics, the mapping of graph features, and boundary conditions, refer to the paper Learning Reduced-Order Models for Cardiovascular Simulations with Graph Neural Networks.
AI surrogate: Dataset, architecture, and experiments
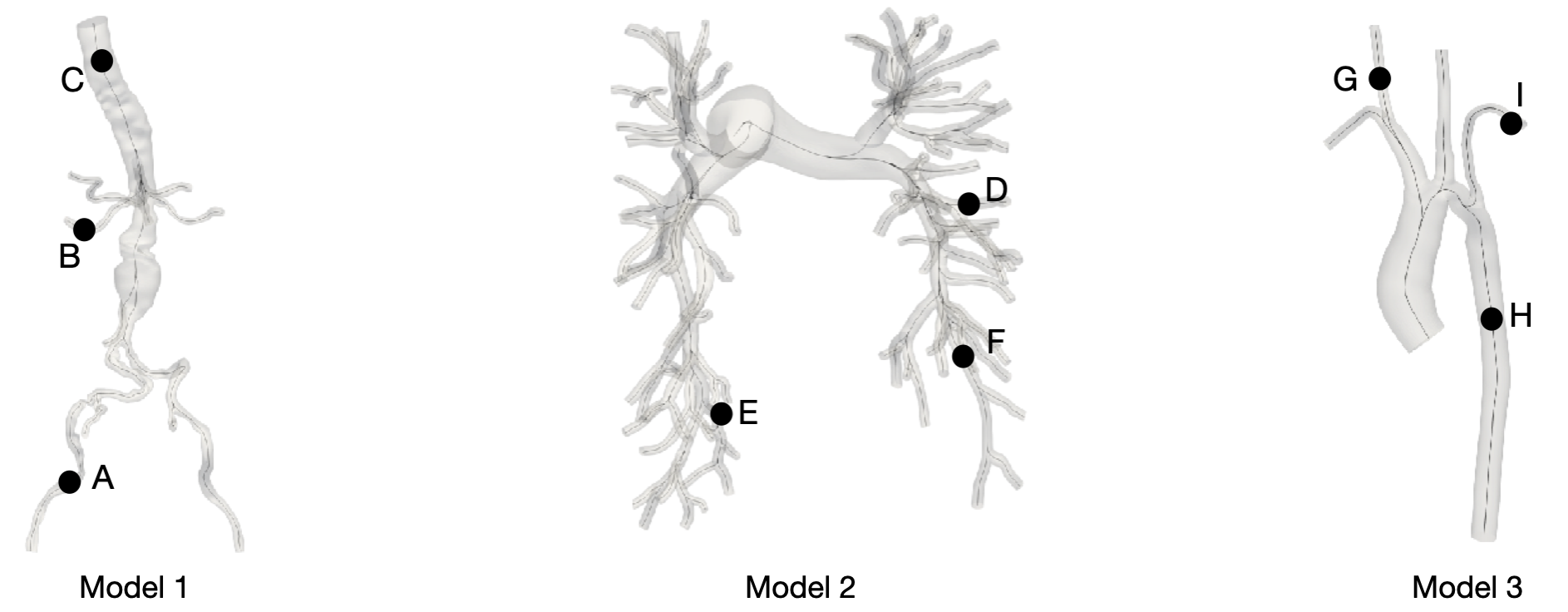
The researchers used models from the Vascular model repository, containing about 252 cardiovascular models. They picked eight models, capturing typically challenging scenarios. Figure 3 shows three of the eight models: an aortofemoral model featuring an aneurysm (Model 1), a healthy pulmonary model (Model 2), and an aorta model affected by coarctation (Model 3). These include features such as multiple junctions (Model 2) or stenoses (Model 3), which can be challenging to handle using 1D physics-based models.
Data was generated using 3D finite-element simulations of the unsteady Navier-Stokes flow. This was done using the SimVascular software suite. The simulation data was then transformed into a 1D centerline representation by averaging the quantities of interest along orthogonal sections at each node.
Simulations were done with different boundary conditions and for each geometry. Two cardiac cycles were simulated using 50 random configurations of boundary conditions to generate the training dataset. You can download the training dataset using the scripts from the NVIDIA PhysicsNeMo GitHub repo.
The MeshGraphNet architecture was chosen to model the system and was modified to suit the cardiovascular simulations. Like the base architecture, the model consists of three components: an encoder, a processor, and a decoder (Figure 2). The encoder transforms the node and edge features into latent representations using a fully connected neural network (FCNN).
Then the processor performs several rounds of message passing along edges, updating both node and edge embeddings. The decoder extracts nodal pressure and flow rate at each node, which is used to update the mesh in an auto-regressive manner. For the exact details of the Meshgraphnet architecture, you can refer to the Learning Mesh-Based Simulation with Graph Networks.
The team ran various experiments to analyze:
- The convergence of the rollout error as a function of the dataset size.
- Sensitivity analysis to evaluate which node and edge features were more important to the accuracy of predicting the flow rate and pressure
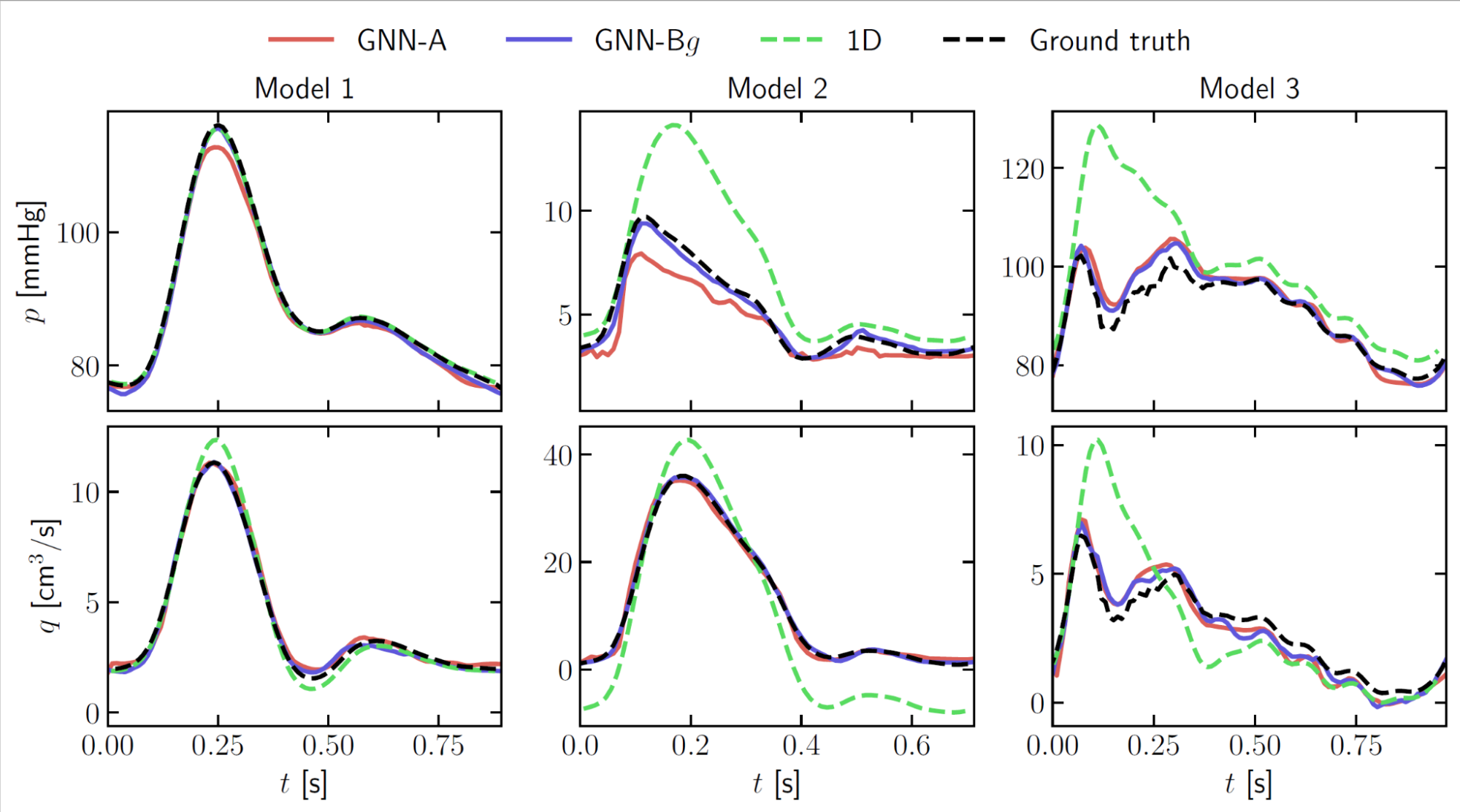
- Direct comparisons with physics-driven one-dimensional models showed superior performance of the GNN, especially when handling complex geometries such as those with many junctions (Figure 4).
- Different approaches to train the algorithm: training networks specific to different cardiovascular regions instead of a single network able to handle different geometries (GNN-A vs GNN-Bg in Figure 4).
The research team is continuing to explore further work related to the generalization of the GNN to more geometries as well as exploring the optimal feature set for improving the performance. Furthermore, they are looking to extend this work to 3D models but also to leverage such physics-ML models into their Simvascular software suite to accelerate simulations.
Using NVIDIA PhysicsNeMo for your research
NVIDIA PhysicsNeMo is an open-source project under the Apache 2.0 license to support the growing physics-ML community. If you are an AI researcher working in the field of physics-informed machine learning and want to get started with NVIDIA PhysicsNeMo go to the PhysicsNeMo GitHub repo.
If you would like to contribute your work to the project, follow the contribution guidelines in the project or reach out to the NVIDIA PhysicsNeMo team.
NVIDIA is celebrating developer contributions across use cases, demonstrating how to build and train physics-ML models using the NVIDIA PhysicsNeMo framework. Equally important is the effort to systematically organize such innovative work in the PhysicsNeMo open-source project for the community and the ecosystem to leverage for their engineering and science surrogate modeling problems.
To learn more about how PhysicsNeMo is being used by the industry, you can refer to the PhysicsNeMo resources page.