Watch the Analysing Cassandra Data using GPUs workshop.
Organizations keep much of their high-speed transactional data in fast NoSQL data stores like Apache Cassandra. Eventually, requirements emerge to obtain analytical insights from this data. Historically, users have leveraged external, massively parallel processing analytics systems like Apache Spark for this purpose. However, today’s analytics ecosystem is quickly embracing AI and ML techniques whose computation relies heavily on GPUs.
In this post, we explore a cutting-edge approach for processing Cassandra SSTables by parsing them directly into GPU device memory using tools from the RAPIDS ecosystem. This enables you to reach insights faster with less initial setup and also make it easy to migrate existing analytics code written in Python.
In this post, I take a quick dive into the RAPIDS project and explore a series of options to make data from Cassandra available for analysis with RAPIDS. Ultimately, I describe the current approach: parsing SSTable files in C++ and converting them into a GPU-friendly format, making the data easier to load into GPU device memory.
To skip the step-by-step journey and try out sstable-to-arrow now, see the next post in this series, Analyzing Cassandra Data using GPUs, Part 2.
What is RAPIDS
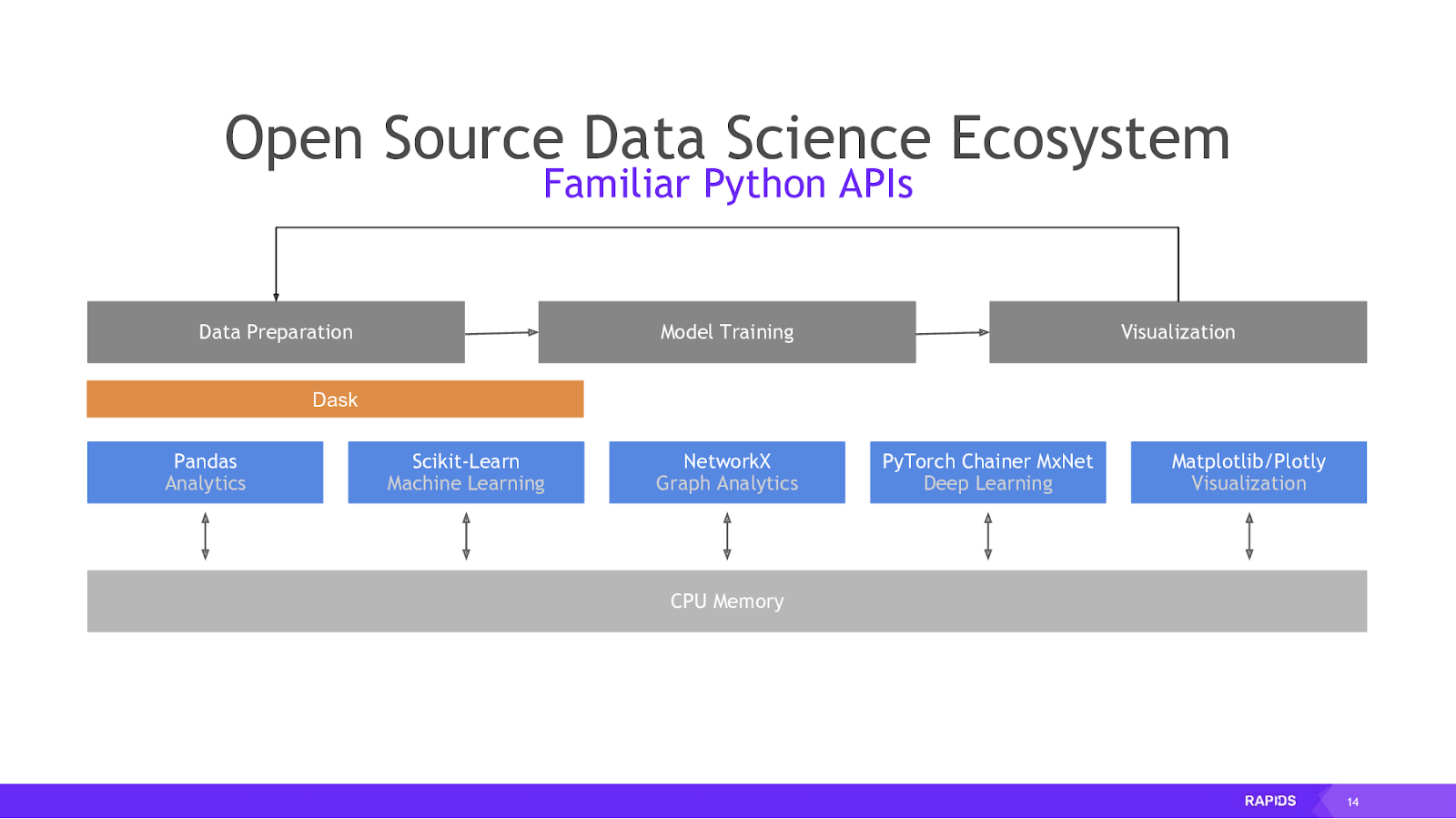
RAPIDS is a suite of open source libraries for doing analytics and data science end-to-end on a GPU. It emerged from CUDA, a developer toolkit developed by NVIDIA to empower you to take advantage of your GPUs.
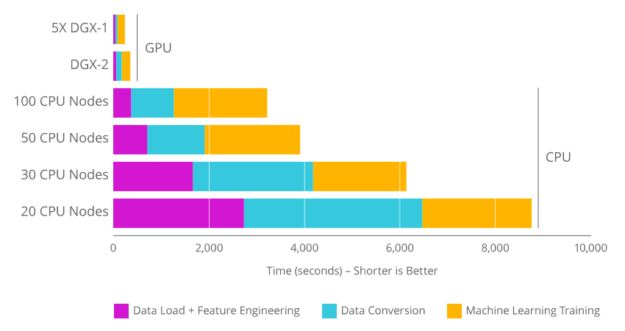
RAPIDS takes common AI / ML APIs like pandas and scikit-learn and makes them available for GPU acceleration. Data science, and particularly machine learning, uses numerous parallel calculations, which makes it better-suited to run on a GPU, which can multitask at a few orders of magnitude higher than current CPUs:
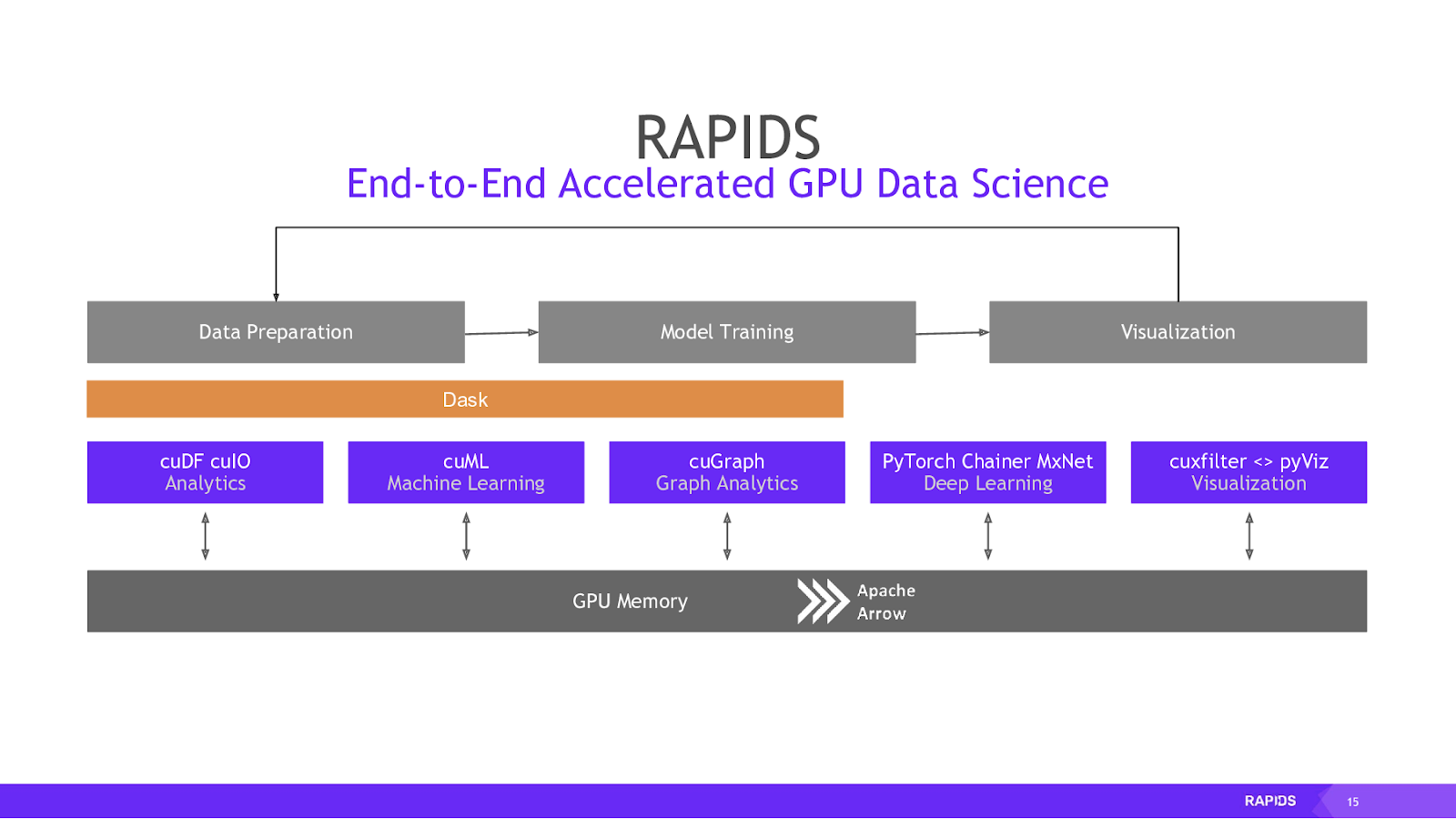
When you get the data on the GPU in the form of a cuDF (essentially the RAPIDS equivalent of a pandas DataFrame), you can interact with it using an almost identical API to the Python libraries that you might be familiar with.
Figure 3 shows Apache Arrow as the underlying memory format. Arrow is based on columns rather than rows, causing faster analytic queries. It also comes with inter-process communication (IPC) mechanism used to transfer an Arrow record batch (a table) between processes. The IPC format is identical to the in-memory format, which eliminates any extra copying or deserialization costs and gets you some extremely fast data access.
The benefits of running analytics on a GPU are clear. All you need is the proper hardware, and then you can migrate your existing data science code to run on the GPU by finding and replacing the names of Python data science libraries with their RAPIDS equivalents.
How do you get Cassandra data onto the GPU?
Over the past few weeks, I have been looking at five different approaches, listed in order of increasing complexity:
- Fetch the data using the Cassandra driver, convert it into a pandas DataFrame, and then turn it into a cuDF.
- Same as the preceding, but skip the pandas step and fetch data using the Cassandra driver directly into Arrow.
- Read SSTables from the disk using Cassandra server code, serialize it using the Arrow IPC stream format, and send it to the client.
- Same as approach 3, but use a custom SSTable parser in C++ instead of using Cassandra code.
- Same as approach 4, but integrate CUDA to speed up table reads while parsing the SSTables.
I give a brief overview of each of these approaches, compare them, and then explain the next steps.
Fetch data using the Cassandra driver
In this approach, you can use existing libraries without having to do too much hacking. Grab the data from the driver, setting session.row_factory to your pandas_factory function to tell the driver how to transform the incoming data into a pandas.DataFrame. Then, call the cudf.DataFrame.from_pandas function to load your data onto the GPU, where you can then use the RAPIDS libraries to run GPU-accelerated analytics.
The following code example requires you to have access to a running Cassandra cluster. For more information, see the DataStax Python Driver documentation. You also must install the required Python libraries with Conda:
BashCopy
conda install -c blazingsql -c rapidsai -c nvidia -c conda-forge -c defaults blazingsql cudf pyarrow pandas numpy cassandra-driver
PythonCopy
from cassandra.cluster import Cluster
from cassandra.auth import PlainTextAuthProvider
import pandas as pd
import pyarrow as pa
import cudf
from blazingsql import BlazingContext
import config
# connect to the Cassandra server in the cloud and configure the session settings
cloud_config= {
'secure_connect_bundle': '/path/to/secure/connect/bundle.zip'
}
auth_provider = PlainTextAuthProvider(user=’your_username_here’, password=’your_password_here’)
cluster = Cluster(cloud=cloud_config, auth_provider=auth_provider)
session = cluster.connect()
def pandas_factory(colnames, rows):
"""Read the data returned by the driver into a pandas DataFrame"""
return pd.DataFrame(rows, columns=colnames)
session.row_factory = pandas_factory
# run the CQL query and get the data
result_set = session.execute("select * from your_keyspace.your_table_name limit 100;")
df = result_set._current_rows # a pandas dataframe with the information
gpu_df = cudf.DataFrame.from_pandas(df) # transform it into memory on the GPU
# do GPU-accelerated operations, such as SQL queries with blazingsql
bc = BlazingContext()
bc.create_table("gpu_table", gpu_df)
bc.describe_table("gpu_table")
result = bc.sql("SELECT * FROM gpu_table")
print(result)
Fetch data using the Cassandra driver directly into Arrow
This step is identical to the previous one, except you can switch out pandas_factory with the following arrow_factory:
PythonCopy
def get_col(col):
rtn = pa.array(col) # automatically detects the type of the array
# for a full implementation, fully check which arrow types want
# to be manually casted for compatibility with cudf
if pa.types.is_decimal(rtn.type):
return rtn.cast('float32')
return rtn
def arrow_factory(colnames, rows):
# convert from the row format passed by
# CQL into the column format of arrow
cols = [get_col(col) for col in zip(*rows)]
table = pa.table({ colnames[i]: cols[i] for i in
range(len(colnames)) })
return table
session.row_factory = arrow_factory
You can then fetch the data and create the cuDF in the same way.
However, both of these two approaches have a major drawback: They rely on querying the existing Cassandra cluster. You don’t want to do this because the read-heavy analytics workload might affect the transactional production workload, where real-time performance is key.
Instead, see if there is a way to get the data directly from the SSTable files on the disk without going through the database. This brings me to the next three approaches.
Read SSTables from the disk using Cassandra server code
Probably the simplest way to read SSTables on disk is to use the existing Cassandra server technologies, namely SSTableLoader. When you have a list of partitions from the SSTable, you can manually transform the data from Java objects into Arrow vectors corresponding to the columns of the table. Then, you can serialize the collection of vectors into the Arrow IPC stream format and then stream it in this format across a socket.
The code here is more complex than the previous two approaches and less developed than the next approach, so I have not included it in this post. Another drawback is that although this approach can run in a separate process or machine than the Cassandra cluster, to use SSTableLoader, you first want to initialize embedded Cassandra in the client process, which takes a considerable amount of time on a cold start.
Use a custom SSTable parser
To avoid initializing Cassandra, we developed our own custom implementation in C++ for parsing the binary data SSTable files. For more information about this approach, see the next post in this series, Analyzing Cassandra Data using GPUs, Part 2.
The guide to the Cassandra storage engine by The Last Pickle helped a lot when deciphering the data format. We decided to use C++ as the language for the parser to anticipate eventually bringing in CUDA and also for low-level control to handle binary data.
Integrate CUDA to speed up table reads
We plan to start working on this approach when the custom parsing implementation becomes more comprehensive. Taking advantage of GPU vectorization should greatly speed up the reading and conversion processes.
Comparison
At the current stage, you are mainly concerned with the time it takes to read the SSTable files. For approaches 1 and 2, you can’t actually measure this time fairly, because the approach relies on additional hardware (the Cassandra cluster) and there are complex caching effects at play within Cassandra itself. However, for approaches 3 and 4, you can perform simple introspection to track how much time the program takes to read the SSTable file from start to finish.
Table 4 shows the results against datasets with 1K, 5K, 10K, 50K, 100K, 500K, and 1M rows of data generated by NoSQLBench:
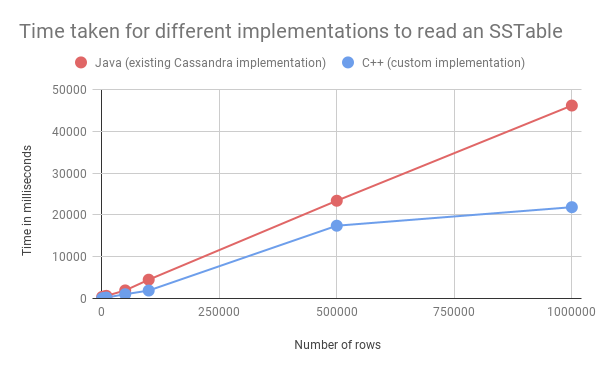
As the graph shows, the custom implementation is slightly faster than the existing Cassandra implementation, even without any additional optimizations such as multithreading.
Conclusion
Given that data access patterns for analytical use cases usually include large scans and often reading entire tables, the most efficient way to get at this data is not through CQL but by getting at SSTables directly.
We implemented a sstable parser in C++ that can do this and convert the data to Apache Arrow so that it can be leveraged by analytics libraries, including the NVIDIA GPU-powered RAPIDS ecosystem. The resulting open-source (Apache 2 licensed) project is called sstable-to-arrow and it is available on GitHub and accessible through Docker Hub as an alpha release.
We will be holding a free online workshop on analyzing Cassandra data with GPUs, which will go deeper into this project with hands-on examples in mid-August!
If you are interested in trying out sstable-to-arrow, look at the next post in this series, Analyzing Cassandra Data using GPUs, Part 2. Feel free to reach out to seb@datastax.com with any feedback or questions.