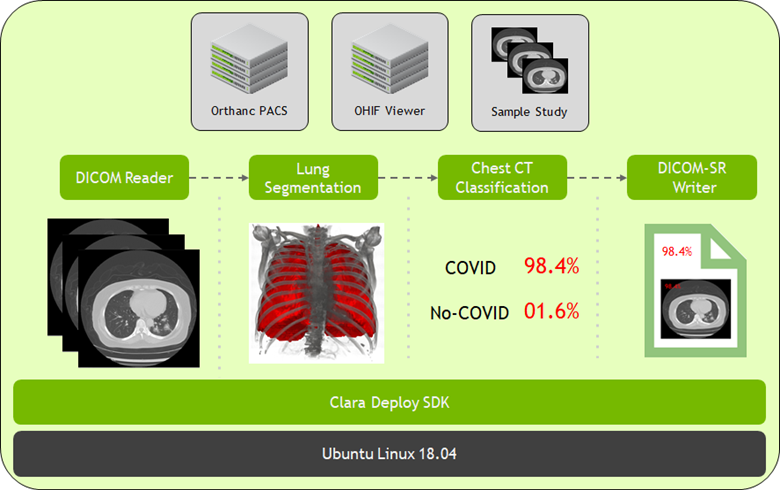
Getting AI up and running in hospitals has never been more important. Until recently, connecting an inference pipeline to perform analysis has had its challenges and limitations. There is a considerable amount of complexity in setting up and maintaining the hardware and software, deployment, configuration, and all workflow steps in an AI inference research pipeline. NVIDIA Clara Deploy, an application framework for running imaging inference pipelines at scale, can be used to streamline this process.
In this post, we demonstrate the NVIDIA Clara Deploy QuickStart Virtual Machine (VM) using a COVID-19 inference pipeline. With the NVIDIA Clara VM and the following scripts, getting started is seamless. NVIDIA has put together a simple three-step process to get your hospital environment ready to deploy inference at scale:
- Deploy the base VM: Use a clean Ubuntu Linux 18.04 instance.
- Run the installation scripts: Download and execute the Clara Deploy Ansible scripts, currently in early access.
- Trigger the pipeline: Open the sample CT study with the embedded web client, run the job using the preinstalled Clara Deploy pipeline, and visualize the inference results.
A successful deployment of any deep learning-based application requires meaningful integration with healthcare information systems. The inference platform that is used to execute deep learning-based workflows need to support industry standards such as DICOM for ingestion of input data and publishing of inference results.
Clara VM is preconfigured to accept data from outside and can be integrated with an external PACS.

Inside the Clara VM
The Clara VM relies on Clara Deploy, a framework for creating AI workflows, using pipelines and operators.
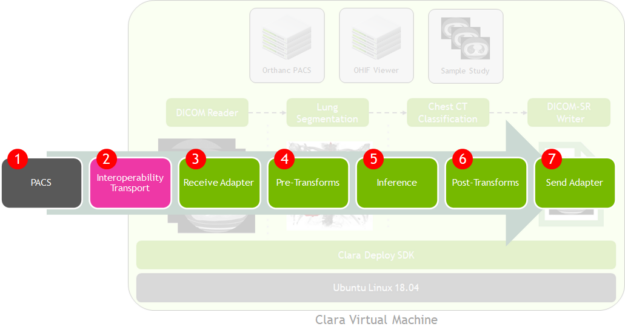
A Clara Deploy instantiation follows the same pattern:
- PACS: This system holds DICOM images acquired from modalities, where you configure a send destination to Clara.
- Interoperability transport: Within the PACS, DICOM images are sent routing rules or through manual pushes.
- Receive adapter: Clara Deploy DICOM Adapter receives the DICOM images.
- Pre-transforms: Clara Deploy converts DICOM imaging data into a format that the inference engine, TRITON, can process.
- Inference: NVIDIA Triton, part of Clara Deploy, performs inference on the imaging study to derive observations on the data.
- Post-transforms: The observations are transformed into a format, like a DICOM Segmentation, Structured Report (DICOM-SR), or HL7 FHIR message.
- Send adapter: Clara Deploy transmits these observation messages to the destination system, such as PACS.
All of this processing happens locally and no processing is performed externally. After it is set up, no connection to the internet is required. It is important that you apply the appropriate security measures that you would apply to any system that has potentially sensitive data.
In addition to Clara Deploy, there are other components included in Clara VM to accelerate the deployment experience: an image repository using Orthanc PACS, a DICOM web viewer with OHIF, and a sample study to demonstrate how the pipeline works.
Deploy the Clara VM
The following instructions walk you through the steps necessary to provision a VM, configure the Clara Deploy installation, and use the included Orthanc repository and and OHIF web viewer to visualize results of the Clara COVID Segmentation pipeline. Clara VM is currently in early access, which requires approval to access the scripts from NGC. Sign up for early access.
To get started, you only need two components from a hardware perspective:
- Infrastructure to deploy a virtual machine on (for example, VMware, or a cloud instance)
- Access to a NVIDIA GPU, such as NVIDIA T4, V100, or A100
The installation scripts assume that you have access to a system with a passthrough T4, 16 vCPU, 32 GB RAM, and a 500-GB disk. If you have something different, some adjustments may need to be tweaked.
Step 1: Deploy the base VM
To streamline this process, you can download a VM ISO with the platform preconfigured, allowing you to begin interacting with the platform immediately.
Deploy a basic Ubuntu OS on the VM instance.
Step 2: Run the installation script
Configuration of the VM is done using Ansible, a tool for repeatably configuring bare metal or virtualized machines. Ansible can be used either locally or from a separate provisioning system. In this case, use a local installation, which requires a few simple prerequisites.
First, you update, install dependencies, add the required repositories, and then install Ansible:
sudo apt update
sudo apt install -y software-properties-common unzip
sudo apt-add-repository --yes --update ppa:ansible/ansible
sudo apt install -y ansibleTo pull the Ansible scripts for installing and configuring the NVIDIA driver and Clara Platform, you must install the NGC CLI and configure it to use your NGC API key. For more information about installation instructions, including generating the NGC API key, see Introduction to the NGC Catalog and Registry CLIs. For this VM, install as follows:
mkdir ~/bin && cd ~/bin wget -O ngccli_reg_linux.zip \ https://ngc.nvidia.com/downloads/ngccli_reg_linux.zip unzip -o ngccli_reg_linux.zip chmod u+x ngc md5sum -c ngc.md5 echo "export PATH=\"\$PATH:$(pwd)\"" >> ~/.bash_profile source ~/.bash_profile ngc config set
The NGC config set command prompts for your NGC API key. Defaults may be accepted for the other configuration options. You can then use the NGC CLI to download the Clara Ansible scripts:
ngc registry resource download-version \ "ea-nvidia-clara/clara/clara_ansible:0.7.3-2011.5" cd clara_ansible_v0.7.3-2011.5 unzip ansible.zip cd playbooks
The playbooks directory contains the Ansible scripts to install and configure the NVIDIA driver and Clara Platform. This installation happens in three steps:
- Install and configure the NVIDIA driver.
- Install the Clara Platform.
- Install and configure the COVID-19 pipeline.
If you already have the NVIDIA driver and Clara Deploy up and running, you can skip the first two steps and start with the pipeline installation.
To configure this for the local system, two configuration changes to the clara_hosts file are required:
- Add the following entry for
localhostunder the[clara_hosts]section (one line):localhost ansible_connection=local ansible_python_interpreter="{{ansible_playbook_python}}" - Add your NGC API key in the last line of the file
First, install the NVIDIA driver:
ansible-playbook -K install-nvidia-driver.yml -i clara_hosts sudo reboot # manual reboot is required because you’re running locally
Then install and configure the Clara Platform:
ansible-playbook -K install-clara.yml -i clara_hosts
The final step is configuring the COVID-19 pipeline and Orthanc/OHIF endpoints.
ansible-playbook -K install-clara-pipeline.yml -i clara_hosts
These Ansible scripts download and install everything needed for Clara Deploy, as well as stage and start the COVID-19 “classification+lesion segmentation” pipeline, listen on a DICOM endpoint, and install Orthanc and OHIF for visualization of results. When the Ansible script is finished, it is ready to accept DICOM to automatically process and send the results into the local Orthanc for visualization in OHIF.
Step 3: Trigger the pipeline
An example COVID study has been registered with Orthanc. To view this study, navigate to the Orthanc interface at http://<IP of the VM>/pacs-admin/app/explorer.html. This prompts you to log in. The Orthanc instance has been configured with username/password orthanc/orthanc. After logging in, you see the main Orthanc interface. To see all available studies, choose All Studies.
On the All Studies screen, you see the example Chest CT COVID-19 study that has been preloaded. Open to see the study details.
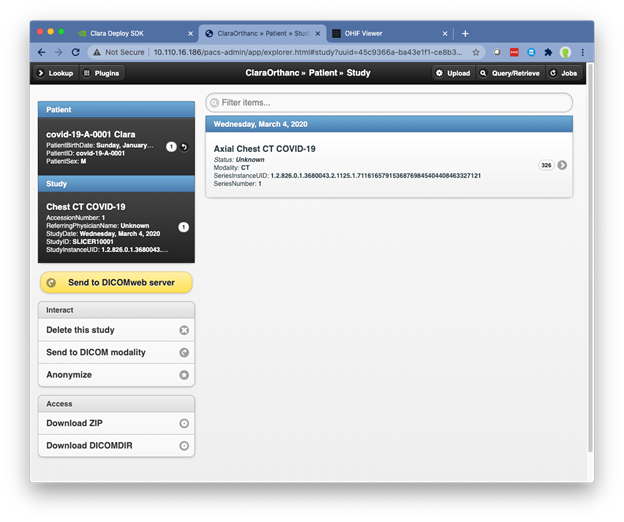
In the left menu, choose Send to DICOM Modality, covid19. This triggers the Clara Deploy pipeline.
Review the results
After the pipeline has completed, refresh this view and you see that the study has been updated with the results of the pipeline, including the following:
- Lung Segmentation
- Lung Lesion Segmentation
- Two DICOM reports
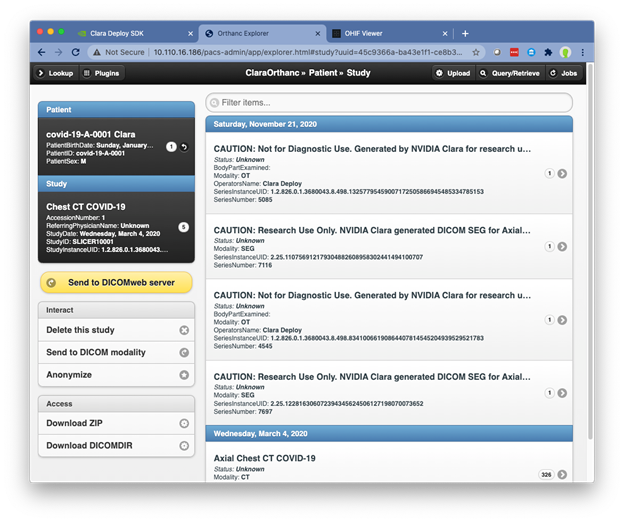
This study can now be visualized in OHIF by navigating to http://<IP address of the VM>. In the main OHIF interface, choose Study List where you see the Chest CT COVID-19 study. Open this study to navigate to the visualization interface. You can view the lung and lesion segmentations along with DICOM reports that detail the results of COVID classification. Figure 6 shows an OHIF layout with panels showing the segmentation as well as the DICOM report with the COVID classification.

Considerations and next steps
When you are sending studies from your environment, only non-multiframe CT studies are accepted. These are denoted with the SOP Class 1.2.840.10008.5.1.4.1.1.2. Secondary captures may not be properly processed.
In the Clara VM for COVID, the Clara pipeline has been preconfigured with DICOM AE titles and source and destination endpoints to interface with Orthanc. To enable this kind of integration, under the hood, the Clara DICOM Adapter provides a set of tools to build this interface between an external PACS, like Orthanc, and the Clara Platform.
To send studies for processing to the Clara VM, perform a CSTORE to the IP address of the Clara VM, on port 4242, with an AE title of “CLARA”. This can be configured inside of a PACS by a PACS administrator by logging into the application and adding a send destination to the Clara VM. After the NVIDIA Clara DICOM Adapter receives the DICOM study, processing can begin.
Extending Clara VM with the COVID-19 Classification Pipeline
For this example, use the Clara Deploy AI COVID-19 Classification Pipeline available on NGC. Following the instructions there, pull and configure the pipeline:
mkdir -p ~/.clara/pipelines && cd ~/.clara/pipelines clara pull pipeline clara_ai_covid19_pipeline cd clara_ai_covid19_pipeline unzip app_covid-19-input_v1.zip -d input sudo unzip app_covid-19-model_v1.zip -d /clara/common/models/ clara create pipeline -p COVID-19-pipeline.yaml
This results in a pipeline with a unique pipeline ID. You can create the DICOM source and destinations for Orthanc and then use this pipeline ID to create a DICOM AE title that is used to trigger the pipeline from Orthanc:
clara dicom create source -a ORTHANC -i 10.110.16.186
clara dicom create destination -a ORTHANC -i 10.110.16.186 \
-n MYPACS -p 4242
pipeid=$(clara list pipelines | grep COVID | awk '{print $2}')
clara dicom create aetitle -a covid19 \
pipeline-COVID-19-pipeline=${pipeid}
The COVID-19 pipeline includes a sample dataset that you can push to Orthanc. This is the dataset used in the preceding Orthanc/OHIF example. To push this dataset to Orthanc, use the DICOM toolkit as follows
storescu -v +sd +r -xb -v -aet DCM4CHEE \ -aec ORTHANC <IP of the VM> 4242 \ ~/.clara/pipelines/clara_ai_covid19_pipeline/input/dcm/
This sample study is now accessible from the Orthanc UI, which can be used to trigger the pipeline using the covid19 AE title (modality), as in the preceding example.
This can be extended further to integrate into the medical imaging ecosystem. Follow the earlier outline to create a new pipeline using one of the example pipelines bundled with Clara Deploy. Following these steps, you can create a unique AE title, push an example dataset to Orthanc, trigger the pipeline, and visualize the results in OHIF.
You can also look more closely at the Clara Platform and pipelines using the Clara console. Install using the Clara CLI:
clara pull -y console clara console start
After the Clara console is started, it is accessible at http://<IP of the VM>:32002. Use the console to view details of the pipelines and jobs that were triggered through the Orthanc interface. Figure 7 shows the Clara console Job view with the DAG for the COVID-19 pipeline.

Figure 7. The Clara Deploy console showing the DAG for the COVID-19 pipeline.
Summary
In this post, we walked you through the steps to go from a base Ubuntu VM to a fully configured Clara Deploy Platform with the COVID-19 segmentation pipeline integrated with a PACS repository and results viewer. This serves as a blueprint for deploying applications within hospital infrastructure to enable Clara Deploy inference at scale.
For more information, request early access to Clara VM or see the Scalable and Modular AI Deployment Powered by NVIDIA Clara Deploy platform whitepaper.