A new study out of the Massachusetts Institute of Technology (MIT) could arm healthcare workers with the information needed to effectively treat COVID-19 patients. Recently published in the Proceedings of the National Academy of Sciences, the research develops a deep learning model that determines the best drug combinations for fighting the virus, despite having relatively limited data.
“Typically, researchers build deep learning models using large existing datasets for things like cancer and cardiovascular disease, but they can’t be used for new illnesses with limited data. Our work shows that we can overcome the data scarcity challenge by incorporating different kinds of biological knowledge,” said lead author Wengong Jin, a postdoctoral associate at the Eric and Wendy Schmidt Center of Broad Institute, and MIT PhD.
With a large number of Americans unvaccinated, breakthrough cases on the rise, and a looming threat of variants, effective drug combinations remain central to mitigating COVID-19. Treatment options such as antiviral drugs, therapeutic monoclonal antibodies, and corticosteroids have proven effective. But there’s little clarity around optimal treatment combinations, while also limiting possible side effects.
Understanding these drug synergies could help patients recover more quickly, increase the likelihood of survival, and reduce pressure on hospital resources.
Training deep learning algorithms to identify drug combinations effective in treating a disease typically requires large datasets. As a new virus with less data—compared to illnesses such as cancer, HIV, or heart disease—COVID-19 poses a greater challenge for model development.
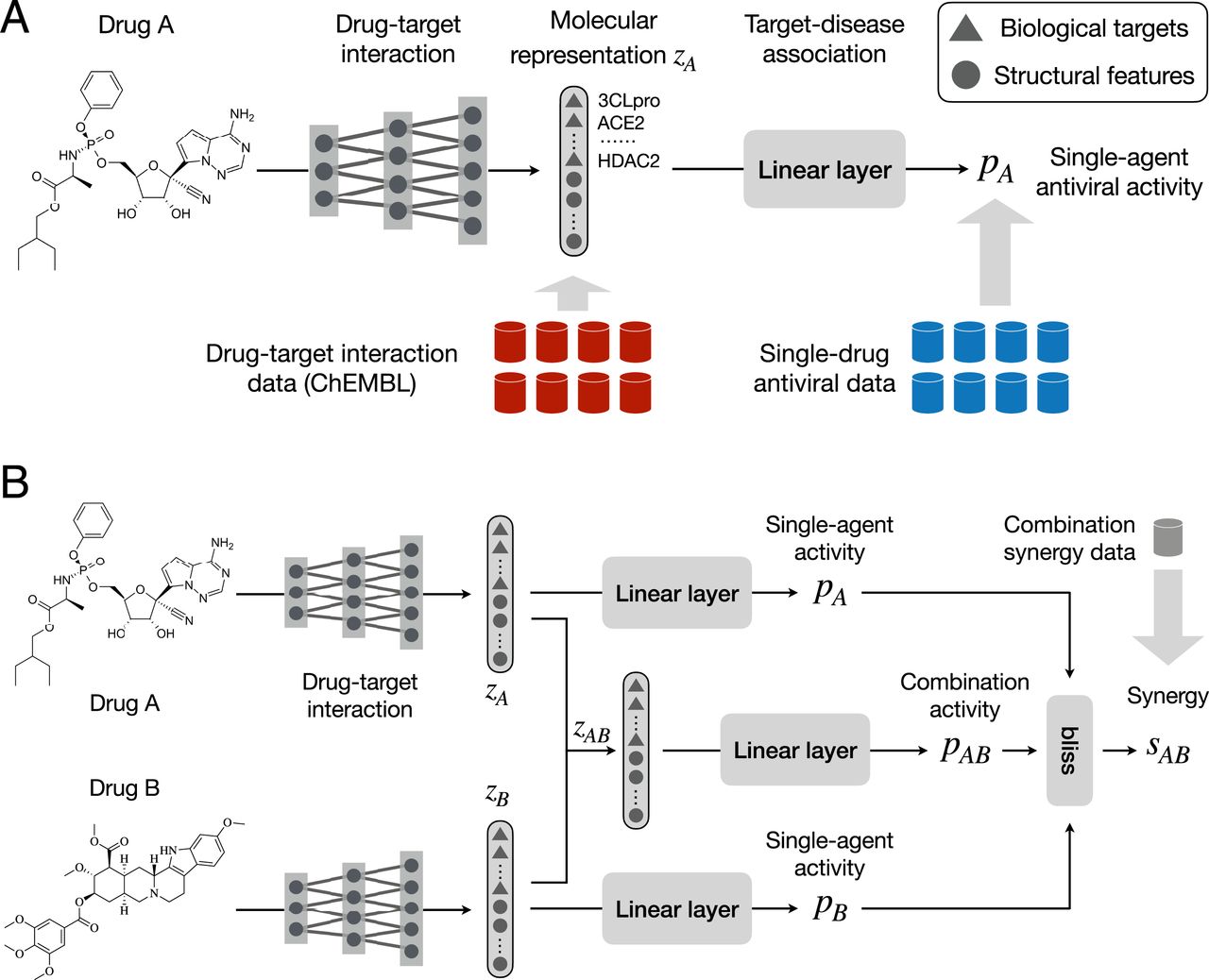
The researchers created a method capable of working with limited data, using a new, two-pronged approach. First, the team trained a neural network to predict whether a drug will bind to a biological target. These targets play a major role in drug therapy by creating a place for a drug to bind and inhibit disease growth. In the case of COVID-19, these targets include enzymes and proteins involved in viral replication.
Based on the molecular structure of a drug and biological targets of the disease, a model also calculates the antiviral effectiveness of a single drug. Based on this information a synergy prediction model combines calculates the potency of drug treatments, determining the most effective combinations.
The neural network models employ an NVIDIA GPU and cuDNN-accelerated deep learning framework to train and process the data. Running 88 different treatment options, the team identified two prime duos for fighting the virus—the antiviral drug remdesivir combined with the hypertension drug reserpine, and remdesivir used alongside IQ-1S (a kinase inhibitor.)
The model can also be applied to other pathogens. According to Jin, the team has been working with the National Institutes of Health, to find drug combinations for treating pancreatic cancer.
The researchers plan to make the code open source sometime this month.
Read the full article in the Proceedings of the National Academy of Sciences >>
Read more >>
