The NVIDIA Clara AGX development kit with the us4R ultrasound development system makes it possible to quickly develop and test a real-time AI processing system for ultrasound imaging. The Clara AGX development kit has an Arm CPU and high-performance RTX 6000 GPU. The us4R teamwork provides ultrasound system designers with the ability to develop, prototype, and test end-to-end software-defined ultrasound systems. Clara AGX is launching the era of software-defined medical instruments with reconfigurable pipelines without changes to the hardware.
The us4us hardware and SDK provide an end-to-end ultrasound algorithm development and RF processing platform, while the high-end Clara AGX GPU enables real-time deep learning and AI image reconstruction and inferencing. With this approach, the whole systems engineering team benefits: beamforming experts can create optimal beam strategies, and AI experts can design and deploy the next generation of real-time algorithms.
This combined hardware and software platform democratizes ultrasound development for both research labs and commercial vendors to develop novel features. No longer are massive capital budgets required to design, prototype, and test functional hardware. Each stage of the device pipeline can be modified. Data acquisition, data processing, image reconstruction, image processing, AI analysis, and visualization are all defined in software and executed in real time with low latency performance. The system is completely configurable and can create new RF transmission waveforms and beamforming algorithms using AI or traditional approaches.
With ultrafast, low latency end-to-end data transfer is possible with NVIDIA ConnectX-6 SmartNIC 100Gb/s ethernet and RDMA data transfer to the GPU. The NVIDIA supercharged GPU can run circles around existing legacy premium-cost systems. It enables improved signal-to-noise, real-time processing of highly advanced and complex algorithms in image reconstruction, denoising, and pipelines.
The GPU has enough headroom for multiple real-time clinical inferencing predictions to run simultaneously, including measurement, operator guidance, image interpretation, tissue and organ identification, advanced visualization, and clinical overlays.
Commercialization of clinical applications of the Clara AGX hardware will be available with medical-grade hardware from third party vendors in a compact and energy-efficient CPU+GPU SoC form-factor similar to that in self-driving automotive applications.
The Clara AGX development kit is a high-end performance workstation built with development of medical applications in mind. The system includes an NVIDIA RTX 6000 GPU, delivering 200+ INT8 AI TOPs and 16.3 FP32 TFLOPS at peak performance, with 24 GB of VRAM. This leaves plenty of headroom for running multiple models. High-bandwidth I/O communication with sensors is possible with the 100G Ethernet Mellanox ConnectX-6 network interface card (NIC).
NVIDIA partners are currently using the Clara AGX development kit to develop ultrasound, endoscopy, and genomics applications.
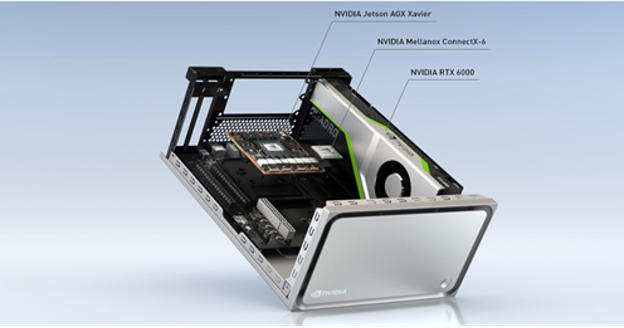

us4R and NVIDIA Clara AGX
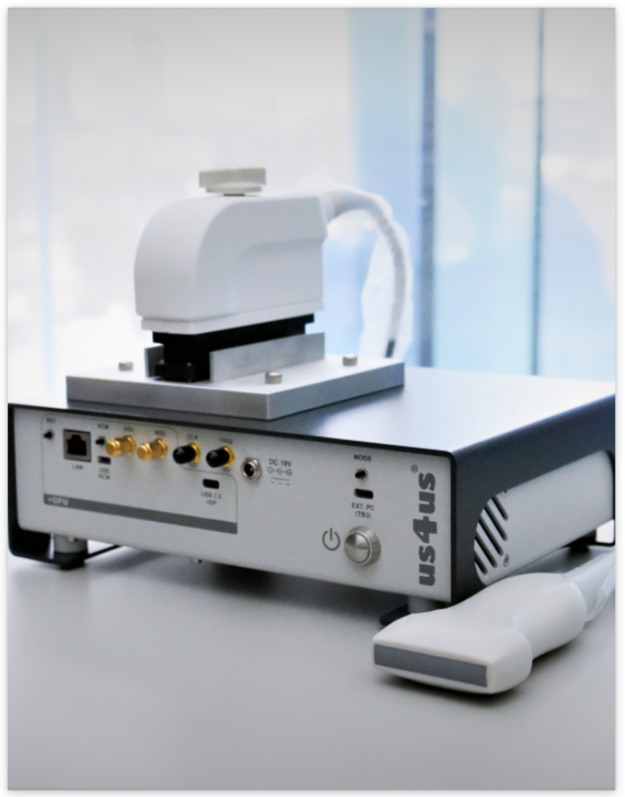

us4us Ltd. offers two systems:
- Advanced us4R with up to 1024 TX / 256 RX channels
- A portable us4R-lite with 256 TX / 64 RX channels
Both use a PCIe streaming architecture for low-latency data transfer and GPU for scalable processing of raw ultrasound echo signals. The us4OEM ultrasound frontend modules support 128TX/32RX analog channels and high throughput 3GB/s, PCIe Gen3 x4 data interface (Figure 2).
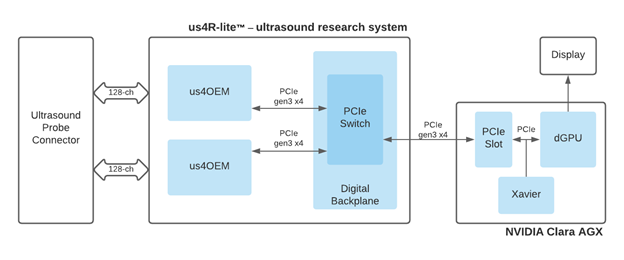
End-to-end, software-defined ultrasound design
The ARRUS package is an SDK for the us4R that provides a high-level hardware abstraction layer enabling systems programming in Python, C++, or MATLAB. The hardware programming is performed by defining RF module including the following:
- Active transmit (TX) probe elements
- Transmission parameters, such as TX voltage, TX waveform, and TX delays
- Receive (RX) aperture and acquisition parameters such as gain, filters, and time-gain compensation
Commonly used TX/RX sequences, such as classical linear scanning, plane wave imaging (PWI) and synthetic transmit aperture (STA) are preconfigured and can be quickly implemented. Custom sequences are configured with user-defined, low-level parameters like TX/RX apertures mask and TX delays.
The ARRUS package also includes a Python implementation of many standard ultrasound processing algorithms for image reconstruction, including raw RF data, RF data preprocessing (data filtering, quadrature demodulation, and so on), beamforming (PWI, STA, and classical schemes), and post-processing of B-mode images.
These algorithms are building blocks used to construct an arbitrary imaging pipeline that can handle the RF data stream produced by the us4R system. GPU accelerated numerical routines are provided by cuPy. DLPack specifies a common in-memory Tensor structure that enables data sharing between machine learning frameworks and GPU processing libraries, while using RDMA no additional overhead is required to copy data between them. The DLPack interface provides access to predefined or user-developed deep learning models in TensorFlow, PyTorch, Chainer, and MXNet.
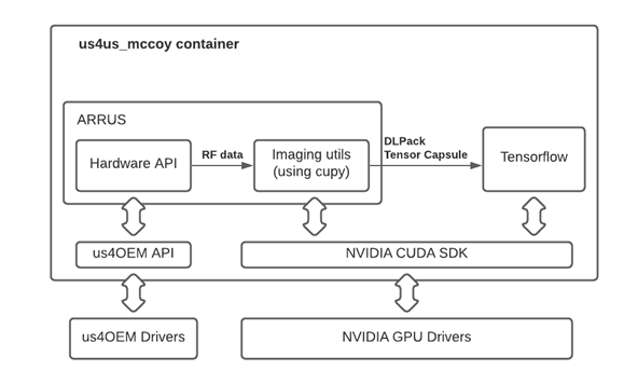
US4US ultrasound demo
By combining the software and hardware stack, you can quickly implement an ultrasound workflow with configurable parameters in less than one page of easy-to-read Python code. In this section, we show you how to use the ARRUS APIs, a us4R-lite platform, and the Clara AGX DevKit to create your own ultrasound imaging pipeline in minutes.
The following code example should work with the proper environment. However, we recommend using the Docker container available directly through NGC. There is an interactive Jupyter notebook available to help guide you through this demo in the container at /us4us_examples/mimicknet-example.ipynb.
Start by importing the relevant libraries, including ARRUS, Numpy, TensorFlow, and CuPy:
# Imports for ARRUS, Numpy, TensorFlow and CuPy import arrus import arrus.session import arrus.utils.imaging import arrus.utils.us4r import numpy as np from arrus.ops.us4r import (Scheme, Pulse, DataBufferSpec) from arrus.utils.imaging import ( Pipeline, Transpose, BandpassFilter, Decimation, QuadratureDemodulation, EnvelopeDetection, LogCompression, Enqueue, RxBeamformingImg, ReconstructLri, Sum, Lambda, Squeeze) from arrus.ops.imaging import ( PwiSequence ) from arrus.utils.us4r import ( RemapToLogicalOrder ) from arrus.utils.gui import ( Display2D ) from utilities import RunForDlPackCapsule, Reshape import TensorFlow as tf import cupy as cp
Next, instantiate the PWI Tx and Rx sequences. You define the parameters for the data that you’re pulling from the US4US Ultrasound system in the PwiSequence function.
seq = PwiSequence( angles=np.linspace(-5, 5, 7)*np.pi/180,# np.asarray([0])*np.pi/180, pulse=Pulse(center_frequency=6e6, n_periods=2, inverse=False), rx_sample_range=(0, 2048), downsampling_factor=2, speed_of_sound=1450, pri=200e-6, sri=20e-3, tgc_start=14, tgc_slope=2e2)
After defining the sequence, load the deep learning model parameters. For this, you have two different deep neural networks options for improving B-mode image output quality, both available for download through NGC.
The NN_Bmode model, from researchers at Stanford, produces despeckled images using a neural network from beamformed low-resolution images (LRI). The LRI is created after a single synthetic aperture transmission; in this case, a single plane wave insonification. A sequence of LRIs can be compounded into a high-resolution image (HRI) by coherently summing them together.
The generative adversarial networks (GANs) model is used to imitate B-mode image post-processing found in commercial ultrasound systems. This algorithm uses the standard delay-and-sum (DAS) reconstruction and B-mode post-processing pipeline with the MimickNet CycleGAN. For more information, see MimickNet, Mimicking Clinical Image Post-Processing Under Black-Box Constraints.
For this example, you load the MimickNet CycleGAN. In addition to loading the weights, you are implementing simple normalize and mimicknet_predict wrapper functions required when you implement the scheme definition in the next step.
# Load MimickNet model weights model = tf.keras.models.load_model(model_weights) model.predict(np.zeros((1, z_size, x_size, 1), dtype=np.float32)) def normalize(img): data = img-cp.min(img) data = data/cp.max(data) return data def mimicknet_predict(capsule): data = tf.experimental.dlpack.from_dlpack(capsule) result = model.predict_on_batch(data) # Compensate a large variance of the image mean brightness. result = result-np.mean(result) result = result-np.min(result) result = result/np.max(result) return result
You can put all of the pieces together using the Scheme function. The Scheme function takes parameters for the tx/rx sequence definition: an output data buffer, the ultrasound device work mode, and a data processing pipeline. These parameters define the workflow for data acquisition, data processing, and displaying the inference results.
The following code example shows the Scheme definition, which includes the sequence, MimickNet preprocessing, and inference wrapper function defined earlier. The placement parameter indicates that the processing pipeline runs on GPU:0, which provides GPU acceleration on the Clara AGX Dev Kit.
scheme = Scheme( tx_rx_sequence=seq, rx_buffer_size=2, output_buffer=DataBufferSpec(type="FIFO", n_elements=4), work_mode="HOST", processing=Pipeline( steps=( ... ReconstructLri(x_grid=x_grid, z_grid=z_grid), # Image preprocessing Lambda(normalize), Reshape(shape=(1, z_size, x_size, 1)), # Deep Learning inference wrapper RunForDlPackCapsule(mimicknet_predict) ... Enqueue(display_input_queue, block=False, ignore_full=True) ), placement="/GPU:0" ) )
Connect to the US4US device, upload your scheme sequence, and start your display queue.
us4r = sess.get_device("/Us4R:0")
us4r.set_hv_voltage(30)
# Upload sequence on the us4r-lite device.
buffer, const_metadata = sess.upload(scheme)
display = Display2D(const_metadata, cmap="gray", value_range=(0.3, 0.9),
title="NNBmode", xlabel="Azimuth (mm)", ylabel="Depth (mm)",
show_colorbar=True, extent=extent)
sess.start_scheme()
display.start(display_input_queue)
print("Display closed, stopping the script.")
Your device now displays the results of the ultrasound imaging pipeline. You can also easily modify this pipeline to implement your own state-of-the-art deep learning algorithms. Figure 5 shows the example output from the demo comparing a conventional delay and sum algorithm (left) and the MimickNet model (right).



The code for this demo is available in a downloadable Docker image through NGC at https://ngc.nvidia.com/catalog/containers/nvidia:clara-agx:agx-us4us-ultrasound.
Conclusion
Clara AGX is launching the era of software-defined medical instruments with re-configurable pipelines without changes to the hardware. Connecting the Clara AGX development kit with the us4R ultrasound development system creates a combination that helps you develop a real-time AI processing system easily and quickly. With the high performance of an RTX 6000 GPU and an Arm CPU, you get the best of the embedded hardware ecosystem to develop your own state-of-the-art, task-specific algorithms.
For more information about the us4R-lite system, contact us4us. The Clara AGX Developer Kit is currently available exclusively for members of the NVIDIA Clara Developer Partner Program.