With the advent of new deep learning approaches based on transformer architecture, natural language processing (NLP) techniques have undergone a revolution in performance and capabilities. Cutting-edge NLP models are becoming the core of modern search engines, voice assistants, chatbots, and more. Modern NLP models can synthesize human-like text and answer questions posed in natural language. As DeepMind research scientist Sebastian Ruder says, NLP’s ImageNet moment has arrived.

While NLP use has grown in mainstream use cases, it still is not widely adopted in healthcare, clinical applications, and scientific research. Part of the reason is that earlier models were trained on Wikipedia and text from literature and did not perform as well on clinical and scientific language. New NLP models such as BioMegatron have been trained by NVIDIA on a large clinical and scientific corpus and achieved excellent performance on common biomedical NLP tasks like named entity recognition (NER), relation extraction (RE) and question answering (QA).
BioMegatron can be used to extract key concepts and relations from biomedical texts and build knowledge graphs that can drive research and discovery. It can also identify clinical terms in clinical speech and text and map them to a standardized ontology to assist in clinical documentation and research. The BioMegatron model was pretrained on an AI cluster of eight DGX-2 for approximately 400 hours, or just about two weeks. This AI cluster is known as a DGX SuperPOD and offers a fast I/O architecture for large AI compute tasks, as well as enough computer power to train the model when new scientific literature is published. Training a larger language model for a longer period, on more clinical and biomedical text using a specialized vocabulary, proved to improve model accuracy.
NLP for pharmaceutical, biotech, and research
For pharmaceutical companies, NLP holds the potential to deliver tremendous value for automating text mining. It could enable pharmaceutical companies to uncover valuable information hidden among troves of unstructured data. Examples of unstructured data are scientific journal articles, physician notes, and medical imaging reports. In the past, unstructured data was manually analyzed and interpreted. Now, cutting-edge NLP techniques make it possible to create large knowledge graphs with ontology mapping from a massive corpus of textual data. These knowledge graphs can then be used for several downstream tasks, starting with powerful semantic search tools.
There are already examples of pharmaceutical companies using NLP for target identification and prioritization, drug repurposing, interpretation of genes and proteins identified by ‘omics experiments, and full patent text mining for new targets. NLP is also being used to extract information on treatment patterns to identify drug switching or discontinuation. NLP can extract and normalize numerical data such as lab values and dosage information, as well as patient-specific details such as a history of disease, problem lists, demographics, social factors, and lifestyle.
NLP for clinicians and hospitals
For clinicians, inputting notes from patient visits is laborious and time-consuming. It’s often completed after hours when they are tired and may have forgotten the entire conversation.
NLP can be incorporated into an automatic speech recognition (ASR) system so that key clinical words can be identified in individual speech or in a conversation. The identified clinical words are then mapped into concepts in a standardized medical ontology. Virtual or in-person patient/clinician interactions are captured and inputted right to medical charts, serving as real-time medical transcription. Clinicians and patients could feel confident that all information is recorded. For clinicians, it frees up time to see more patients or spend more time with patients. A medical speech recognition system could also help with voice-activated queries about a specific part of a patient’s medical history during a visit. “It’s a key enabler for telemedicine and other tools for documentation,” commented Dr. Mona Flores, NVIDIA Global Head of Medical AI. “The doctor can concentrate on the patient, instead of taking notes.”
A clinical NLP AI model can also help with computer-assisted coding, automated registry reporting, data mining, and clinical documentation improvement. This reduces the documentation burden for doctors and hospitals, giving them more time to focus on patients. Figure 1 shows an example of how clinical entities can be extracted using BioMegatron and mapped to ontologies, from the Fast and Accurate Clinical Named Entity Recognition and Concept Mapping from Conversations paper.
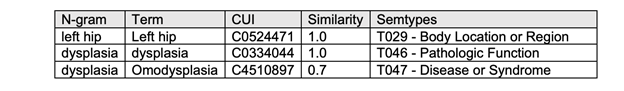
BioMegatron overview
BioMegatron is a state-of-the-art (SOTA) language model for biomedical and clinical NLP developed at NVIDIA. Recent work has demonstrated that larger language models dramatically advance NLP applications such as QA, dialog systems, summarization, and article completion. BioMegatron is the largest biomedical transformer-based language model ever trained. It is up to 3.5x the size of BERT, with 345 million, 800 million, and 1.2 billion parameter variants. It was trained on 6.1 billion words from PubMed, a repository of abstracts and full text journal articles on biomedical topics.
BioMegatron is based on NVIDIA NLP model Megatron-LM, which swaps the position of the layer normalization and the residual connection in the model architecture (similar to GPT-2 architecture). This allows the models to continue to improve as they are scaled up.
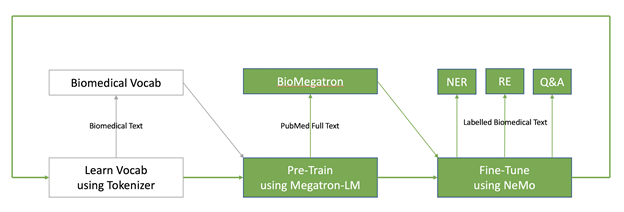
Modern NLP models follow a two-step paradigm of pretraining followed by fine-tuning. Pretraining is done on a large corpus of text (PubMed) in an unsupervised manner, producing a scientific language model (BioMegatron). This language model is then tweaked for a variety of downstream NLP applications like NER, RE, and QA. In the case of domain-specific language models, there’s an additional first step of selecting a good vocabulary to train the language model. In our experience with building BioMegatron, the choice of vocabulary has a big impact on the performance of the downstream NLP models.
The pretraining process is the most computationally intensive step. It involves a significant amount of hyperparameter tuning. As models get larger, they can start bumping up against memory constraints. Model-parallel training is a way to overcome this by splitting the model parameters across multiple GPUs.
The Megatron-LM model provides a simple and efficient model-parallel approach to training transformer models, reaching 76% scaling efficiency on 512 GPUs compared to a fast, single-GPU baseline. For more information, see the Megatron-LM software training recipes or you can download BioMegatron from NVIDIA Clara NLP NGC and get started right away. NVIDIA Clara NLP NGC is a collection of models and resources that supports NLP in healthcare and life sciences.
Fine-tuning was performed using NeMo, which is an open-source toolkit for conversational AI. Using NeMo, BioMegatron produces SOTA performance when fine-tuned for a variety of common NLP tasks like NER, RE, and QA. This work was accepted by the 2020 Conference on Empirical Methods in Natural Language Processing (EMNLP 2020), and is now available on arXiv, BioMegatron: Larger Biomedical Domain Language Model.
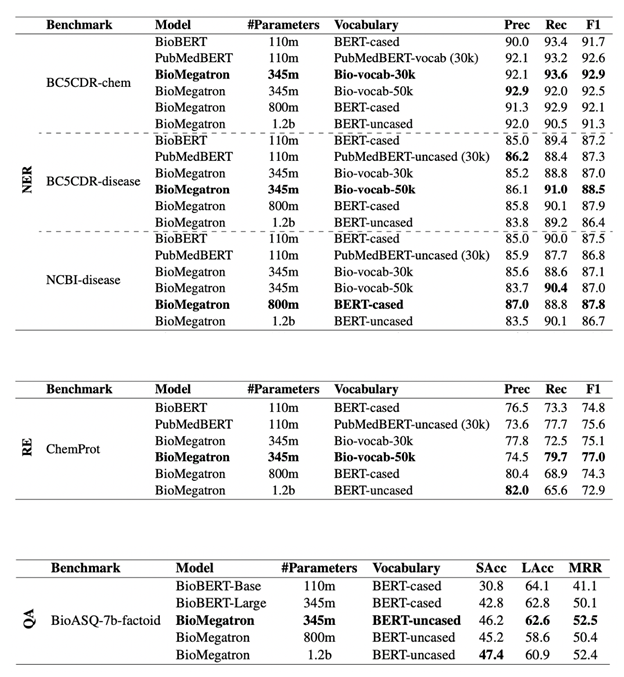
The secret to the BioMegatron SOTA performance is a combination of training a bigger model with a large corpus of text, for a longer time, using scientific biomedical vocabulary and extensive hyperparameter tuning. This was made possible by highly efficient training pipelines and the compute power of a DGX SuperPOD.
Improving BioMegatron
BioMegatron was trained on publicly available biomedical and scientific texts. If you’re in possession of a large corpus of clinical or biomedical texts, like pharma and hospitals, you can train BioMegatron further for even better performance on your specific data.
In the fast-moving world of biomedical and scientific research, new terminologies, drug names, and vocabularies are constantly being discovered and introduced into scientific literature. As a result, consistent training of a language model is essential for optimal accuracy. Get started with BioMegatron for free and test it on your biomedical data or pretrain it further with your data. Download the BioMegatron model and supporting tools.
