NVIDIA Parabricks expands the NVIDIA emphasis on solving omics challenges with deep learning and continues accelerating genomics instruments. NVIDIA Parabricks v4.3.1, released at The European Society of Human Genetics (ESHG), introduces new functionality for variant calling in somatic data and upgrades to the latest versions of industry-leading tools. This release follows the recent Parabricks v4.3 release announced at NVIDIA GTC 2024.
Here are the new features:
- Support for Google’s DeepSomatic in short-read sequencing
- Upgraded versions of DeepVariant (1.6.1) and Minimap2 (v2.26)
- Benchmarks from the previous Parabricks 4.3 release

Variant calling for genomic analysis
Variant calling is a critical component of high-throughput sequencing in genomic analysis. It enables scientists to identify variants of whole genomes, exomes, and gene panels for both germline and somatic workflows, leading to a better understanding of disease and potential treatment.
However, variant calling is an incredibly time-intensive and laborious process that requires significant computational resources. It’s true particularly when looking at whole-genome sequencing. Sequence alignment and variant detection alone require enough bandwidth to align studied sequences to reference genomes and then detect variations like insertions or deletions.
As a result, specialized algorithms and tools have been developed to expedite variant calling and enable researchers to perform its critical steps faster and with higher accuracy.
DeepVariant for germline data
One of the most popular tools for variant calling is DeepVariant, a deep-learning–based variant caller developed by Google that can detect a wide range of variants with high accuracy and scale to effectively analyze large datasets. It is particularly valuable for reducing false positives and detecting variants that are often missed by traditional variant callers. Plus, it is open source, making it accessible to anyone who wants to use it.
Germline variants, also known as germline mutations, occur in reproductive cells and are inherited from either parent. DeepVariant is already available in NVIDIA Parabricks for GPU-accelerated germline variant calling and is now upgraded to version 1.6.1 in the latest Parabricks 4.3.1 release.
DeepSomatic for somatic data
Somatic variants, or somatic mutations, occur after conception and affect non-reproductive cells (egg or sperm cells). Unlike germline variants, somatic variants are not hereditary and happen randomly.
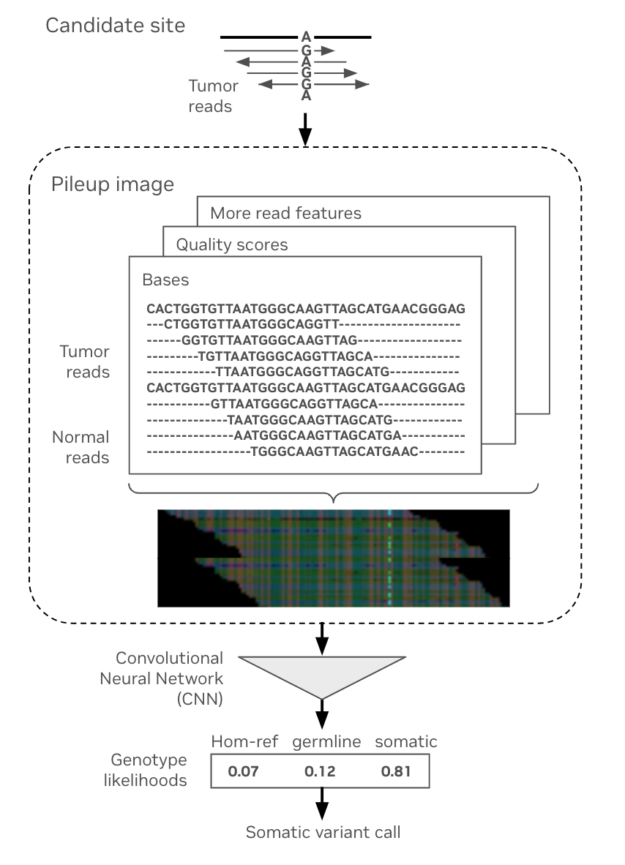
DeepSomatic is the DeepVariant equivalent for somatic data. In the same way that DeepVariant is the deep-learning–based equivalent of GATKs HaplotypeCaller for germline calling, DeepSomatic is the deep-learning–based equivalent of GATKs Mutect2 for somatic calling.
DeepSomatic shares similarities with its germline counterpart DeepVariant, including higher accuracy variant calling and open source availability. However, it is built specifically for somatic data. With the latest 4.3.1 release, Parabricks now supports DeepSomatic for short-read sequencing and harnesses the power of GPU-acceleration for somatic variant calling.
“High-accuracy deep-learning tools like DeepSomatic are critical to advancing genomics research and deepening our understanding of somatic mutations,” explains Francisco Garcia, Ph.D., senior vice president of Informatics at Element Biosciences. “Combined with Element’s high-quality Q50-enabled UltraQ sequencing, they provide a powerful solution for analyzing high-depth cancer genomes. We are excited to use the GPU-accelerated tool made available in the latest release of Parabricks.”

Minimap2 v2.26 upgrade in NVIDIA Parabricks
Minimap2 is a popular tool used to align long-read sequences against a large reference database. Even with insertions, deletions, and inversions, Minimap2 effectively aligns long sequencing. This makes it particularly useful for sequencing platforms, such as PacBio, that analyze long-read sequencing data.
The latest Minimap v2.26 upgrade includes improved splice alignment for RNA sequencing data and improved integration with long-read instrument providers. PacBio, one of these long-read instrument providers, built pbmm2 as a wrapper of Minimap2 for mapping long-read sequencing data produced from their sequencing platforms.
“This latest release of Parabricks includes the same version of minimap2 used by the pbmm2 read aligner from PacBio,” explains Aaron Wegner, senior director of product management at PacBio. ”I am excited to see partners like NVIDIA continuing to make it easier and faster to analyze HiFi long reads from our game-changing Revio system.”

The latest benchmarks for Minimap2 performance show a runtime of 28.7 minutes with four L4 GPUs and 25.6 minutes with only two NVIDIA H100 GPUs (based on a 35x whole genome sequenced from PacBio data).
Parabricks benchmarks
In addition to new features and upgrades for each release, NVIDIA works to continuously improve benchmark performance across instruments, tools, and GPUs.
Table 1 outlines the latest benchmarks from the previous Parabricks v4.3 release on the most popular NVIDIA GPUs for the fastest speed (H100) and lowest cost per sample (L4).
| H100 Fastest Speed | L4 Lowest Cost Per Sample | ||||
| 2 GPU | 4 GPU | 2 GPU | 4 GPU | 8 GPU | |
| FQ2BAM | 17.18 | 9.88 | 47.35 | 21.77 | 13.60 |
| BWA-Meth | 27.43 | 15.12 | 77.35 | 39.77 | 22.47 |
| DeepVariant | 9.67 | 5.82 | 23.48 | 13.10 | 7.8 |
| HaplotypeCaller | 10.57 | 4.90 | 12.00 | 7.73 | 4.27 |
| Mutect2 | 25.80 | 13.60 | 55.8 | 32.50 | 17.5 |
30x whole genome sequenced for FQ2BAM, BWA-Meth, DeepVariant, and Haplotype Caller with Illumina data.
50x tumor-normal whole genome sequenced for Mutect2 with Illumina data.
Get started
With the latest 4.3.1 release, scientists and researchers conducting cancer sequencing can now access DeepSomatic for short-read sequencing. Parabricks 4.3.1 accelerates the deep-learning–based approach from Google by powering an easy-to-use, accelerated version for somatic variant calling.
To download and get started today, see NVIDIA Parabricks.